Spectral pre-processing for 1D NMR
The spectral preprocessing for 1D NMR (1H & 13C) can be automatically applied in case where the input raw data are FID. The term pre-processing designates here the transformation of the NMR spectrum from time domain to frequency domain, including the phase correction and the fast fourier-transform (FFT).(For more details, see Processing methods)
Here, we suggest a reference that could be read with great profit:
James Keeler (2010) Understanding NMR Spectroscopy, 2nd Edition, Ed Wiley
In NMRProcFlow, you can adjust some parameters. Just click on the 'Parameters' button to bring up the window of the parameters to be adjusted, as shown bellow:
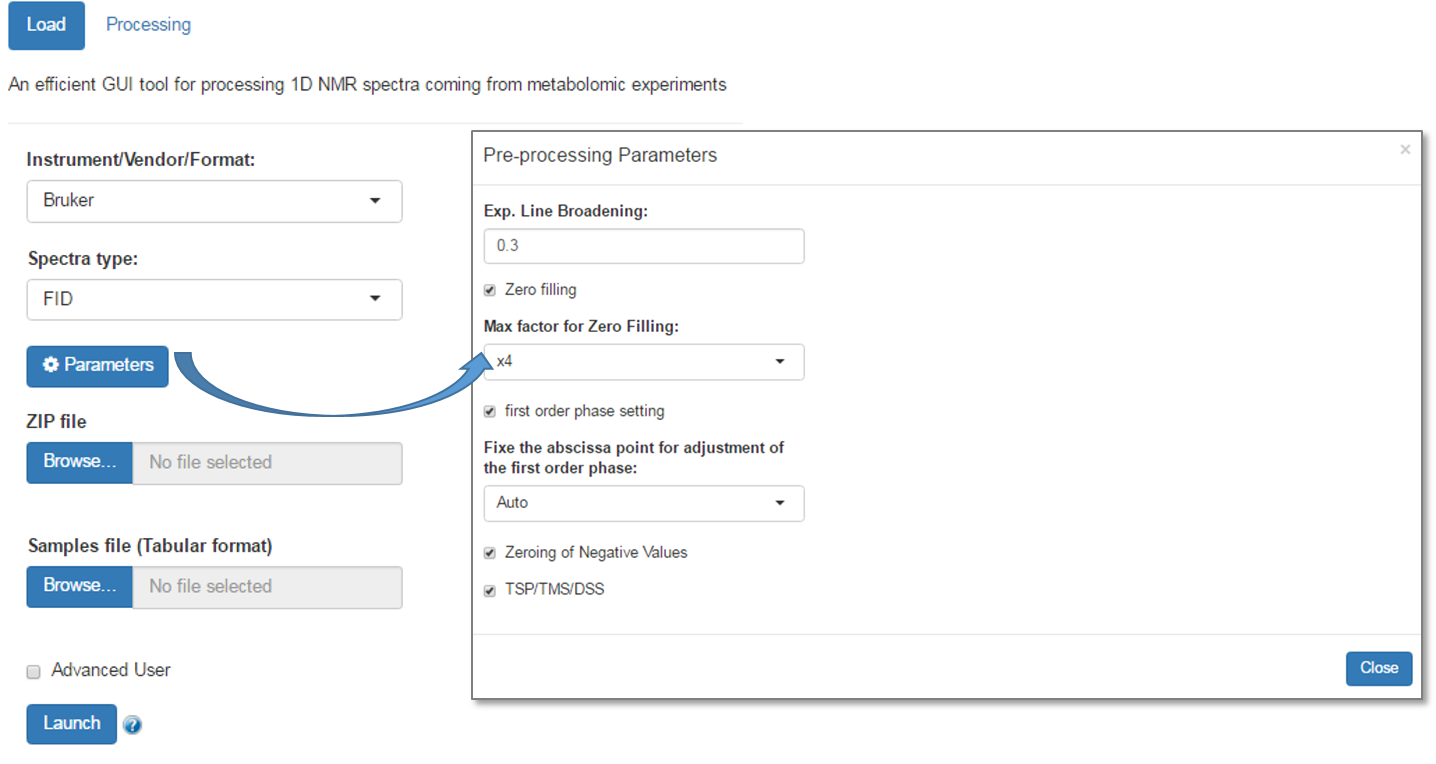
- Line Broadening: Apodization is based on a Line Broadening (i.e an exponential) applied on the fid in order to improve the signal-noise ratio. You can modify the parameter (LB). Zero means no apodization. If necessary, by playing very slightly on the LB parameter, sometimes this may greatly improve the phase correction. Warning: higher the LB value, poorer the resolution.
- Zero filling: This consists of adding zeros at the end of the FID signal such as the resulting size is an even multiple of the initial size. Contrary of an apodization, this has less effect on the spectra resolution, while improving the signal-to-noise ratio.
- First order phase can be adjusted (by default) or not. In case of the First order phase needs to be adjusted, the abscissa point (α) used to set the first order phase can be automatically fixed, otherwise you can choose a value in the dropbox list.
Note: You can optimize the parameters to apply on your own raw spectra from the small application online at the URL https://pmb-bordeaux.fr/nmrspec/.